
Roberto Toro
Institut Pasteur, France

The challenge will be carried out on the RAMP platform. It enables competition and collaboration on data-science problems, using the Python language. To start "hacking", a starting kit is available. It provides a simple working example which can be expanded to more advanced solutions.
Autism spectrum disorder (ASD) is a severe psychiatric disorder that affects 1 in 166 children.
There is evidence that ASD is reflected in individuals brain networks and anatomy. Yet, it remains unclear how systematic these effects are, and how large is their predictive remain unclear. The large cohort assembled here can bring some anwsers. Predicting autism from brain imaging will provide biomarkers and shed some light on the mechanisms of the pathology.
The goal of the competition is to predict the diagnostic status from brain imaging data in the hidden test set. The data of the test set are on a server, hidden from participants. Prediction is done by submitting Python code that will be first trained by the server on training images, and then applied to predict on the hidden test set.
The best performers will be awarded prizes at the end of the competitive period, based on the metrics used during the challenge: 1st - 3000 €, 2nd - 2000 €, 3rd - 1000 €, from 4th to 10th- 500 €.
This challenge is made of 2 phases:
Anyone can participate remotely. In addition, two events will be organized as follows:
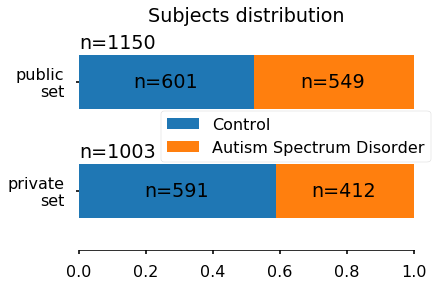
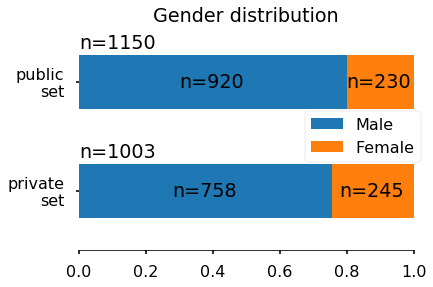
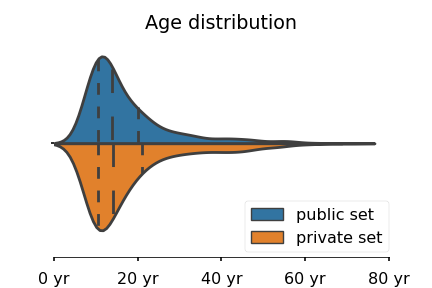
For each subject, a set of structural features have been extracted and stored in a CSV file. Each row corresponds to a subject and each column to a specific features. The features extracted are:
Each subject also comes with fMRI signals extracted on different brain parcellations and atlases, and a set of confound signals. The brain atlases and parcellations are:
The time series for each subject are stored in a CSV file. Each row represent a scan while each column a region defined by the brain parcellation or atlas used.
The data are openly downloadable and a Jupyter notebook provides a starting kit. It explain: (i) how to read data, (ii) how design, train, and test a model, (iii) how to submit it on the competition platform.
Refer to the starting kit for more information.
The group of Neuroanatomy of the Unit of Human Genetics and Cognitive Functions is interested in the development and evolution of brain anatomy. We are particularly interested in Autism Spectrum Disorders.

The Robert Debre Hospital investigates different fields such as genetic, brain imaging, neurocognitive science to identify new biomarkers as well as cell models for new therapeutical targets.

The Paris-Saclay CDS aims at developing methods and tools so as to be capable of analyzing large amounts of data and extracting useful information from them for multidisciplinary sciences.

Ingenieurs et Scientifiques de France (IESF) is the unique representative of the engineers and scientists social corps. The IESF is a major actor of the scientific, technological, and economic debates.

Institut Pasteur, France

Institut Pasteur, France

Institut Pasteur, France

Max Planck Institute, Germany

Inria Parietal, France

Inria Parietal, France

LAL, France

Paris-Saclay Center for Data Science, France

Paris-Saclay Center for Data Science, France

Paris-Saclay Center for Data Science, France
We report the most frequent questions related to the IMPAC challenge. You can visit the GitHub issue tracker for additional information.
You need to register on the RAMP platform and sign up for the IMPAC challenge.
Unfortunately, we cannot share the raw imaging data. This is mostly for privacy and ethical reasons.
Any open-source library available in conda
or pip can be added to the platform. Please open a
ticket on
the GitHub
issue tracker.